Table of Contents
HDF5 Multifile
Maintainer
Overview
The “HDF5 Multifile” project is inspired by the ability of a parallel file system to write several files simultaneously to different storage devices. This feature holds an enormous potential for high performance I/O, but to use this feature efficiently an application must be able to distribute the data uniformly on a large group of storage devices. In this context HDF5 is a special case. It was developed for large datasets and is often used on parallel file systems. Obviously it could benefit from parallel I/O in a large extent.
The goal of our multifile patch is to increase HDF5 I/O performance on a parallel file system. The basic idea is to split HDF5 datasets into several portions and write them to different files. This process isn't fully transparent to the programmer. An application developer has still to take care of the partitioning of datasets and the rest is done automatically by our patch.
Detailed description
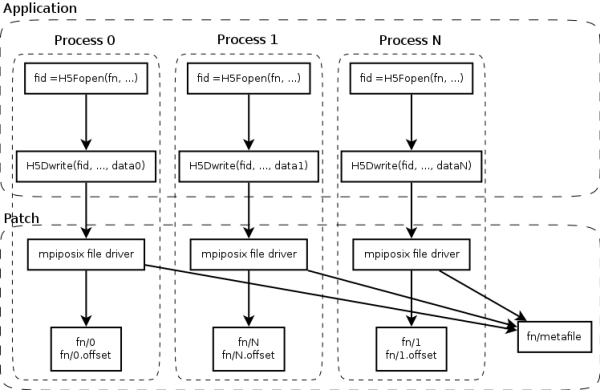
Our work introduces a new HDF5 file driver named “mpiposix”. It implements the HDF5 file driver interface and can be used in same manner as other HDF5 drivers. In the patched HDF5 library it will be set to default. It is up to programmer is to prepare the application for parallel I/O. Typically, that means to assign parts of a dataset to the processes and let them write the data simultaneously to the files.
When a file is created, the patched HDF5 library will create a directory with the given name instead. In this directory, a small file “metafile” will be created that basically contains only the number of process files present. This is only written once by process 0.
Each process creates two files within the directory: a file with its process number as a name and one with the suffix “.offset” appended to its process number. The first holds all the data that the process writes, the second holds the ranges within the reconstructed file where each block of data should end up. This allows us to use purely appending write operations while writing in parallel.
When the file is subsequently opened for reading, one process reads all the individual files, creates a new file, and replays the recorded writes into this file. Of course, this operation takes longer than the original write, but it is much faster than a parallel write to a shared file would have been. After the file is successfully reconstructed, the directory is replaced by the reconstructed file, which is just a conventional HDF5 file. This file is opened and can be read normally.
We have tried a number of different reconstruction algorithms, the code for three of them are available with the patch above, you can select them by commenting/uncommenting #define statements at the top of “src/H5FDmpio_multiple.c”. The default is to read the entire data at once, close the input files, then write the output file in one go. Another, less memory demanding, available option is to use mmap() to read the input, a strategy which should give good performance on a well designed OS. We have not made it the default, though, since there are important systems which fail to deliver good performance with mmap().
The reconstruction can also be parallelized by uncommenting another #define at the top of “src/H5FDmpio_multiple.c”, however this leads to bad performance in all cases.
Performance
Currently, our implementation is limited to the parallel write access. The read operations are still done sequentially. Moreover, the data files need to be merge before first read access. After that you probably get a typical HDF5 read performance.
It is to be expected that maximum I/O performance can be achieved only, when writing to different files on different storage devices by independent processes. Our “POSIX” benchmark was designed in this way and it is our goal to achieve the save results for “HDF5 Multifile” tests.
Performance snapshots
- Test system: Mistral
- Benchmark tool: IOR
- Option details can be found in the IOR user quide
| Number of Nodes | Tasks Per Node | HDF5 collective IO | HDF5 big chunk | HDF5 | POSIX | HDF5 Multifile | POSIX Multifile |
|---|---|---|---|---|---|---|---|
| [MiB/s] | [MiB/s] | [MiB/s] | [MiB/s] | [MiB/s] | [MiB/s] | ||
| 10 | 1 | 815 | 4625 | 805 | 9035 | 9793 | 9308 |
| 10 | 2 | 580 | 4971 | 741 | 17652 | 18210 | 17579 |
| 10 | 4 | 741 | 4685 | 1200 | 30181 | 20672 | 31688 |
| 10 | 6 | 424 | 2498 | 508 | 24330 | 25303 | 27320 |
| 10 | 8 | 618 | 2985 | 403 | 23946 | 24109 | 26872 |
| 10 | 10 | 699 | 2862 | 785 | 29000 | 23497 | 28225 |
| 10 | 12 | 835 | 3160 | 880 | 24642 | 24387 | 24573 |
| 100 | 1 | 807 | 26101 | 865 | 57178 | 70135 | 80430 |
| 100 | 2 | 536 | 17835 | 449 | 73947 | 69233 | 87095 |
| 100 | 4 | 572 | 14390 | 490 | 66982 | 85109 | 92546 |
| 100 | 6 | 1109 | 3390 | 1207 | 76719 | 82949 | 95199 |
| 100 | 8 | 1336 | 3559 | 1222 | 67602 | 84927 | 80288 |
| 100 | 10 | 833 | 4180 | 1090 | 71053 | 80499 | 80691 |
| 100 | 12 | 1259 | 4220 | 1300 | 65997 | 85381 | 85987 |
- HDF5 collective IO: Collective IO is activated in IOR and ROMIO optimizations are used.
- blockSize = 10 MiB, transferSize = 10 MiB
- HDF5 big chunk: One large data set is written in one rush to the file.
- segmentCount = 1
- HDF5: Only ROMIO optimizations are used.
- blockSize = 10 MiB, transferSize = 10 MiB
- POSIX: Independent IO using posix interface.
- blockSize = 10 MiB, transferSize = 10 MiB
- POSIX Multifile: Independent IO using posix interface and patched library.
- blockSize = 10 MiB, transferSize = 10 MiB
- HDF5 Multifile: Patched library without optimizations.
- blockSize = 10 MiB, transferSize = 10 MiB)
- ROMIO optimizations
striping_unit = 1048576 striping_factor = 64 romio_lustre_start_iodevice = 0 direct_read = false direct_write = false romio_lustre_co_ratio = 1 romio_lustre_coll_threshold = 0 romio_lustre_ds_in_coll = disable cb_buffer_size = 16777216 romio_cb_read = automatic romio_cb_write = automatic cb_nodes = 10 romio_no_indep_rw = false romio_cb_pfr = disable romio_cb_fr_types = aar romio_cb_fr_alignment = 1 romio_cb_ds_threshold = 0 romio_cb_alltoall = automatic ind_rd_buffer_size = 4194304 ind_wr_buffer_size = 524288 romio_ds_read = automatic romio_ds_write = disable cb_config_list = *:1
Modules (on Mistral only)
The HDF5 Multifile libraries are available on our HPC system “Mistral” and can be easily loaded using the module command:
- Export module path:
export MODULEPATH=$MODULEPATH:/work/ku0598/k202107/software/modules
- Load HFD5 Multifile libraries:
module load betke/hdf5/1.8.16-multifile module load betke/netcdf/4.4.0-multifile
Download
- Changelog:
- Default file driver: mpiposix
- Optimized access to the *.offset files
- H5Tools support
- Bugfixes and code cleanup
Installation
- Download and extract HDF5-1.8.16
- Download and extract the patch to the HDF5 directory
- Change to the HDF5 directory
- Apply the patch
patch -p1 < hdf5-1.8.16-multifile.patch
- Configure, make and install HDF5 in the usual way
Usage
HDF5-C-API
You can use the multifile driver by creating HDF5 files with
hid_t plist_id = H5Pcreate(H5P_FILE_ACCESS); H5Pset_fapl_mpiposix(plist_id, MPI_COMM_WORLD, false); hid_t fileId = H5Fcreate("path/to/file/to/create.h5", H5F_ACC_TRUNC, H5P_DEFAULT, plist_id);
After that they can be read using this sequence
hid_t plist_id = H5Pcreate(H5P_FILE_ACCESS); H5Pset_fapl_mpiposix(plist_id, MPI_COMM_WORLD, false); hid_t fileId = H5Fopen("path/to/file/to/read.h5", H5F_ACC_RDONLY, plist_id);
Example
- Example source code:
- Compilation:
mpicc -o write-multifile write-multifile.c -I"$H5DIR/include/" -L"$H5DIR/lib" -lhdf5
- Running many copies of the program creates a folder “write-multifile.h5” that contains several files.
mpiexec -n 4 ./write-multifile
- The data files are named with the corresponding process numbers. The information in the “*.offset” is needed to merge the data files to an HDF5 file. The metafile stores the number of files and the size of the data.
find ./multifile.h5/ ./multifile.h5/ ./multifile.h5/0 ./multifile.h5/0.offset ./multifile.h5/1 ./multifile.h5/1.offset ./multifile.h5/2 ./multifile.h5/2.offset ./multifile.h5/3 ./multifile.h5/3.offset ./multifile.h5/metafile
H5Tools
H5Tools accept folder name as parameter, when using together with the mpiposix driver. You must ensure that the folder contains a valid set of mpiposix files.
Suppose data.h5 is a folder in the current directory. Both calls are possible:
h5dump data.h5/
h5dump data.h5
Future work
Since we know that HDF5GROUP is working on a new HDF5 implementation (HDF5-VOL ), we change our priorities in its favour. The most important reason for that is probably the support of the third party plugins. Using the new plugin interface, the idea of HDF5-Multifile can be more easily implemented and maintained. The plugin is development in the scope of the ESIWACE project, now. We transferred our knowledge and discontinued the development of the legacy HDF5-Multifile.
You can checkout the HD5-VOL code from the official repository:
svn checkout http://svn.hdfgroup.uiuc.edu/hdf5/features/vol